Review written by Sara Geraghty (QCB, G3)
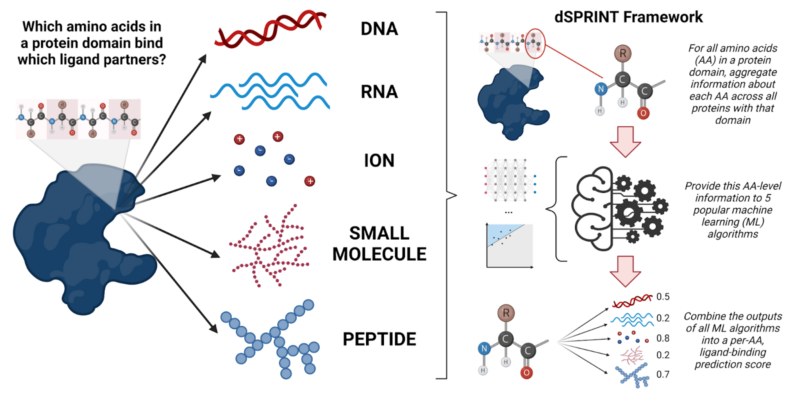
Proteins are ubiquitous in our cells. Currently, it’s estimated that the human body contains between 80,000 and 400,000 protein molecules, all busily performing the many tasks that keep your cells running smoothly and ultimately go into making you. Not only do proteins form the structural framework of your cells, but they also protect your body against foreign pathogens, help digest your food, and send and transmit signals around your body. However, they don’t do this in isolation: proteins are constantly working hand-in-hand with other proteins and molecules in your body, like your DNA, RNA, small molecules, and ions. When your DNA is replicated, or an ion is actively transported, the proteins doing the job need to recognize those molecules (and often bind to them) in order to carry out their task properly. This fundamental process of a protein binding to a partner molecule, or ligand, is critically important in biological processes ranging from development to cancer. And yet, we know surprisingly little about what proteins bind what ligands, and where — knowledge that is key in understanding, and possibly manipulating, the inner workings of cells.
Continue reading