Review by Abigail Stanton (MOL, G1)
Living things are composed of an intricate set of chemical machinery, each piece refined over billions of years of evolution to perform the tasks required to grow, reproduce, act, and react. A principal challenge of biochemistry is understanding how each microscopic gear (or protein) works within the dynamic context of a larger machine (the cell and, eventually, the organism as a whole). To dissect these complex pathways, researchers need ways to interact with the cell. They need tools that act like molecular tweezers to remove pieces, to change them, and to turn mechanisms on and off. As our understanding of each component grows, our biochemical toolbox expands, allowing for even more biological discoveries, which in turn allows for the development of ever more sophisticated tools.
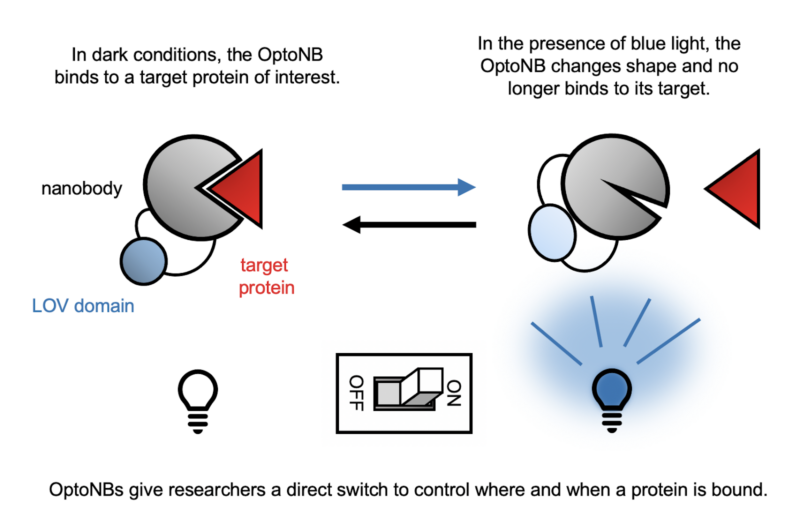
A recent collaboration between the labs of Dr. Jared Toettcher (Molecular Biology) and Dr. José Avalos (Chemical and Biological Engineering) combines two such biochemical tools to control activities in the cell with a literal flick of a switch: nanobodies and optogenetics. Nanobodies are a small form of antibodies produced by camelids such as camels, llamas, and alpacas. They can bind to a specific protein target with high affinity, and they are easily expressed in a range of organisms from bacteria to mammals. Optogenetics is a technique that makes use of proteins that undergo a shape change in response to light. For example, a section of the plant protein phototropin called the light-oxygen-voltage-sensing domain (LOV2) changes shape when exposed to blue light, creating a chemical message that allows the plant to grow towards the sun. The work of the Toettcher and Avalos labs, led by postdoctoral scholar Agnieszka Gil, takes advantage of the shape-shifting ability of the LOV2 domain by inserting it into nanobodies. The addition of LOV2 causes the nanobody to undergo a light-dependent shape change that disrupts its ability to bind to its target, giving the researcher direct control over when and where in the cell the nanobody binds.
To test the feasibility of these optogenetic nanobodies (OptoNBs), the authors made a standard human cell line produce the OptoNB along with a fluorescent tag that would allow them to track its location within the cell. They also had the same cells make the target of this OptoNB, mCherry, with a tag that anchored it to the membrane, the outermost layer of the cell. When the nanobody bound to mCherry, its fluorescent signal would be found on the membrane. Conversely, when binding was disrupted, the nanobody would be found floating freely in the interior of the cell. After testing candidates with the LOV2 domain inserted in ten different locations on the nanobody, the research team found two sites that changed their level of binding in response to blue light. One performed as expected: blue light caused the OptoNB to change shape and lose some ability to bind its target. Surprisingly, the other construct demonstrated the opposite pattern: in this case, light actually increased the level of binding. While mCherry was used for convenience of testing, the same design approach can be easily applied to target any protein for which there is a nanobody available. However, tests with additional nanobodies revealed that the effect of these insertions varied in strength and in some cases, whether light induced binding or dissociation. Therefore, while this approach is flexible for a broad range of targets, novel OptoNBs still require some testing for the optimal location to insert the LOV2 domain.
After demonstrating that OptoNBs could successfully undergo light-dependent binding or dissociation, the authors went on to test how this new tool could be applied to biologically relevant questions. The Toettcher lab is particularly interested in understanding the Erk pathway, a cell signaling pathway that causes cells to divide and helps determine what kind of cell they’ll become during development. By attaching a component of the Erk pathway called SOScat to their mCherry OptoNB, Gil et. al could bring this signal to the membrane, turning the Erk pathway on and off with blue light. This kind of temporal control will allow the Toettcher lab to further probe the dynamics of signaling: not just how a cell responds when a signalling pathway is turned on or off but also how the cell responds to a constant versus oscillating signal. Gil et al. also created an OptoNB targeted to the actin cytoskeleton, the network of protein filaments that give cells their shape and stability. With this construct, the authors demonstrated that light-dependent dissociation could be toggled in a rapid and reversible manner, with the shift between bound and unbound state taking around two minutes. They also demonstrated that by illuminating only one side of the cell, dissociation could be targeted to just that side while the other side remained bound. This subcellular control over binding in the cell could be particularly useful for studies of asymmetrical cell behaviors like polarization or migration.
Gil et al. also demonstrated some potential applications for OptoNB outside of the cellular context. For example, by attaching the OptoNB to small beads made out of agarose, a type of polymer gel used for biochemical applications, mCherry can be pulled from solution to the surface of the bead using light. This system could provide an easy and reversible platform to deplete a target protein from a solution, allowing researchers to determine which components are necessary for a given biochemical activity. One could also potentially use this system to purify a target without the need to use a protein tag that might disrupt the protein’s natural function. However, binding studies using versions of the nanobody that mimic its shape in the light and dark states indicate that the light-induced shape change does not completely get rid of all binding. Instead, it changes the affinity between the nanobody and its target by around four- to sevenfold. This change in affinity is sufficient for signalling applications in the cell, where an initial signal is amplified by a signaling cascade. However, for these applications that take place outside of the cell, a more dramatic shift in binding affinity may be required, and more optimization may be necessary.
This recent work from the Toettcher and Avalos labs opens the door for many exciting applications for expanding our understanding of the inner workings of the cell through direct control of pathways of interest. A key strength of this approach is its versatility. Nanobodies can be produced for a broad range of targets, any of which can be converted to OptoNBs with the insertion of a LOV2 domain. Gil et al.’s research demonstrates that these OptoNBs can be attached to other proteins to direct where they go within the cell, but nanobody binding itself can also sometimes inhibit or activate a target protein. This means that use of OptoNBs will allow researchers to manipulate protein-protein interactions, gene expression, and cell signalling at a specific location and time, all by simply turning a light on and off. As our understanding of the complex machinery of life grows, so too will our ability to make it work for us. Tools like OptoNBs and the many discoveries they enable will also allow us to make new advancements in biochemical engineering, designing new therapeutics and creating cellular factories that make products we need like chemical reagents, drugs, and biofuels.
This original article was published in Nature Communications on August 13, 2020. Please follow this link to view the full version.